Pessimism of the Intellect, Optimism of the Will Favorite posts | Manifold podcast | Twitter: @hsu_steve
Friday, December 31, 2021
Happy New Year 2022!
Friday, December 24, 2021
Peace on Earth, Good Will to Men 2021
When asked what I want for Christmas, I reply: Peace On Earth, Good Will To Men :-)
No one ever seems to recognize that this comes from the Bible (Luke 2.14).
Linus said it best in A Charlie Brown Christmas:
And there were in the same country shepherds abiding in the field, keeping watch over their flock by night.
And, lo, the angel of the Lord came upon them, and the glory of the Lord shone round about them: and they were sore afraid.
And the angel said unto them, Fear not: for, behold, I bring you good tidings of great joy, which shall be to all people.
For unto you is born this day in the city of David a Saviour, which is Christ the Lord.
And this shall be a sign unto you; Ye shall find the babe wrapped in swaddling clothes, lying in a manger.
And suddenly there was with the angel a multitude of the heavenly host praising God, and saying,
Glory to God in the highest, and on earth peace, good will toward men.
Merry Christmas!
The first baby conceived from an embryo screened with Genomic Prediction preimplantation genetic testing for polygenic risk scores (PGT-P) was born in mid-2020.
It is a great honor to co-author a paper with Simon Fishel, the last surviving member of the team that produced the first IVF baby (Louise Brown) in 1978. His mentors and collaborators were Robert Edwards (Nobel Prize 2010) and Patrick Steptoe (passed before 2010). ...
Today millions of babies are produced through IVF. In most developed countries roughly 3-5 percent of all births are through IVF, and in Denmark the fraction is about 10 percent! But when the technology was first introduced with the birth of Louise Brown in 1978, the pioneering scientists had to overcome significant resistance.
There may be an alternate universe in which IVF was not allowed to develop, and those millions of children were never born.
Wikipedia: ...During these controversial early years of IVF, Fishel and his colleagues received extensive opposition from critics both outside of and within the medical and scientific communities, including a civil writ for murder.[16] Fishel has since stated that "the whole establishment was outraged" by their early work and that people thought that he was "potentially a mad scientist".[17]
I predict that within 5 years the use of polygenic risk scores will become common in health systems (i.e., for adults) and in IVF. Reasonable people will wonder why the technology was ever controversial at all, just as in the case of IVF.
And the angel said unto them, Fear not: for, behold, I bring you good tidings of great joy, which shall be to all people.
Mary was born in the twenties, when the tests were new and still primitive. Her mother had frozen a dozen eggs, from which came Mary and her sister Elizabeth. Mary had her father's long frame, brown eyes, and friendly demeanor. She was clever, but Elizabeth was the really brainy one. Both were healthy and strong and free from inherited disease. All this her parents knew from the tests -- performed on DNA taken from a few cells of each embryo. The reports came via email, from GP Inc., by way of the fertility doctor. Dad used to joke that Mary and Elizabeth were the pick of the litter, but never mentioned what happened to the other fertilized eggs.
Now Mary and Joe were ready for their first child. The choices were dizzying. Fortunately, Elizabeth had been through the same process just the year before, and referred them to her genetic engineer, a friend from Harvard. Joe was a bit reluctant about bleeding edge edits, but Mary had a feeling the GP engineer was right -- their son had the potential to be truly special, with just the right tweaks ...
Friday, December 17, 2021
Macroscopic Superposition States: entanglement of a macroscopic living organism (tardigrade) with a superconducting qubit
Entanglement between superconducting qubits and a tardigrade
https://arxiv.org/pdf/2112.07978.pdf
K. S. Lee et al.
Quantum and biological systems are seldom discussed together as they seemingly demand opposing conditions. Life is complex, "hot and wet" whereas quantum objects are small, cold and well controlled. Here, we overcome this barrier with a tardigrade -- a microscopic multicellular organism known to tolerate extreme physiochemical conditions via a latent state of life known as cryptobiosis. We observe coupling between the animal in cryptobiosis and a superconducting quantum bit and prepare a highly entangled state between this combined system and another qubit. The tardigrade itself is shown to be entangled with the remaining subsystems. The animal is then observed to return to its active form after 420 hours at sub 10 mK temperatures and pressure of 6×10−6 mbar, setting a new record for the conditions that a complex form of life can survive.
From the paper:
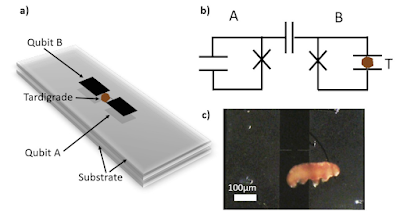
In our experiments, we use specimens of a Danish population of Ramazzottius varieornatus Bertolani and Kinchin, 1993 (Eutardigrada, Ramazzottiidae). The species belongs to phylum Tardigrada comprising of microscopic invertebrate animals with an adult length of 50-1200 µm [12]. Importantly, many tardigrades show extraordinary survival capabilities [13] and selected species have previously been exposed to extremely low temperatures of 50 mK [14] and low Earth orbit pressures of 10−19 mbar [15]. Their survival in these extreme conditions is possible thanks to a latent state of life known as cryptobiosis [2, 13]. Cryptobiosis can be induced by various extreme physicochemical conditions, including freezing and desiccation. Specifically, during desiccation, tardigrades reduce volume and contract into an ametabolic state, known as a “tun”. Revival is achieved by reintroducing the tardigrade into liquid water at atmospheric pressure. In the current experiments, we used dessicated R. varieornatus tuns with a length of 100-150 µm. Active adult specimens have a length of 200-450 µm. The revival process typically takes several minutes.
We place a tardigrade tun on a superconducting transmon qubit and observe coupling between the qubit and the tardigrade tun via a shift in the resonance frequency of the new qubit-tardigrade system. This joint qubit-tardigrade system is then entangled with a second superconducting qubit. We reconstruct the density matrix of this coupled system experimentally via quantum state tomography. Finally, the tardigrade is removed from the superconducting qubit and reintroduced to atmospheric pressure and room temperature. We observe the resumption of its active metabolic state in water.
... a microscopic model where the charges inside the tardigrade are represented as effective harmonic oscillators that couple to the electric field of the qubit via the dipole mechanism... [This theoretical analysis results in the B0 T0 , B1 T1 system where T0 T1 are effective qubits formed of tardigrade internal degrees of freedom.]
... We applied 16 different combinations of one-qubit gates on qubit A and dressed states of the joint qubit B-tardigrade system. We then jointly readout the state of both qubits using the cavity ...
If the creature above (a tardigrade arthropod) can be placed in a superposition state, will you accept that you probably can be as well? And once you admit this, will you accept that you probably actually DO exist in a superposition state already?
It may be disturbing to learn that we live in a huge quantum multiverse, but was it not also disturbing for Galileo's contemporaries to learn that we live on a giant rotating sphere, hurtling through space at 30 kilometers per second? E pur si muove!

Macroscopic Superpositions in Isolated Systems
R. Buniy and S. Hsu
arXiv:2011.11661, to appear in Foundations of Physics
For any choice of initial state and weak assumptions about the Hamiltonian, large isolated quantum systems undergoing Schrodinger evolution spend most of their time in macroscopic superposition states. The result follows from von Neumann's 1929 Quantum Ergodic Theorem. As a specific example, we consider a box containing a solid ball and some gas molecules. Regardless of the initial state, the system will evolve into a quantum superposition of states with the ball in macroscopically different positions. Thus, despite their seeming fragility, macroscopic superposition states are ubiquitous consequences of quantum evolution. We discuss the connection to many worlds quantum mechanics.
... Quantum technology has made great strides over the past two decades and physicists are now able to construct and manipulate systems that were once in the realm of thought experiments. One particularly fascinating avenue of inquiry is the fuzzy border between quantum and classical physics. In the past, a clear delineation could be made in terms of size: tiny objects such as photons and electrons inhabit the quantum world whereas large objects such as billiard balls obey classical physics.
Over the past decade, physicists have been pushing the limits of what is quantum using drum-like mechanical resonators measuring around 10 microns across. Unlike electrons or photons, these drumheads are macroscopic objects that are manufactured using standard micromachining techniques and appear as solid as billiard balls in electron microscope images (see figure). Yet despite the resonators’ tangible nature, researchers have been able to observe their quantum properties, for example, by putting a device into its quantum ground state as Teufel and colleagues did in 2017.
This year, teams led by Teufel and Kotler and independently by Sillanpää went a step further, becoming the first to quantum-mechanically entangle two such drumheads. The two groups generated their entanglement in different ways. While the Aalto/Canberra team used a specially chosen resonant frequency to eliminate noise in the system that could have disturbed the entangled state, the NIST group’s entanglement resembled a two-qubit gate in which the form of the entangled state depends on the initial states of the drumheads. ...
Monday, December 13, 2021
Quantum Hair and Black Hole Information
Quantum Hair and Black Hole Information
https://arxiv.org/abs/2112.05171
Xavier Calmet, Stephen D.H. Hsu
It has been shown that the quantum state of the graviton field outside a black hole horizon carries information about the internal state of the hole. We explain how this allows unitary evaporation: the final radiation state is a complex superposition which depends linearly on the initial black hole state. Under time reversal, the radiation state evolves back to the original black hole quantum state. Formulations of the information paradox on a fixed semiclassical geometry describe only a small subset of the evaporation Hilbert space, and do not exclude overall unitarity.This is the sequel to our earlier paper Quantum Hair from Gravity in which we first showed that the quantum state of the graviton field outside the black hole is determined by the quantum state of the interior.
Our results have important consequences for black hole information: they allow us to examine deviations from the semiclassical approximation used to calculate Hawking radiation and they show explicitly that the quantum spacetime of black hole evaporation is a complex superposition state.
Do Black Holes Destroy Information?
https://arxiv.org/abs/hep-th/9209058
John Preskill
I review the information loss paradox that was first formulated by Hawking, and discuss possible ways of resolving it. All proposed solutions have serious drawbacks. I conclude that the information loss paradox may well presage a revolution in fundamental physics.
Lessons from the Information Paradox
https://arxiv.org/abs/2012.05770
Suvrat Raju
Abstract: We review recent progress on the information paradox. We explain why exponentially small correlations in the radiation emitted by a black hole are sufficient to resolve the original paradox put forward by Hawking. We then describe a refinement of the paradox that makes essential reference to the black-hole interior. This analysis leads to a broadly-applicable physical principle: in a theory of quantum gravity, a copy of all the information on a Cauchy slice is also available near the boundary of the slice. This principle can be made precise and established — under weak assumptions, and using only low-energy techniques — in asymptotically global AdS and in four dimensional asymptotically flat spacetime. When applied to black holes, this principle tells us that the exterior of the black hole always retains a complete copy of the information in the interior. We show that accounting for this redundancy provides a resolution of the information paradox for evaporating black holes ...
Friday, December 10, 2021
Elizabeth Carr: First US IVF baby and Genomic Prediction patient advocate (The Sunday Times podcast)
Genomic Prediction’s Elizabeth Carr: “Scoring embryos”
The Sunday Times’ tech correspondent Danny Fortson brings on Elizabeth Carr, America’s first baby conceived by in-vitro fertilization and patient advocate at Genomic Prediction, to talk about the new era of pre-natal screening (5:45), the dawn of in-vitro fertilization (8:40), the technology’s acceptance (12:10), what Genomic Prediction does (13:40), scoring embryos (16:30), the slippery slope (19:20), selecting for smarts (24:15), the cost (25:00), and the future of conception (28:30). PLUS Dan Benjamin, bio economist at UCLA, comes on to talk about why he and others raised the alarm about polygenic scoring (30:20), drawing the line between prevention and enhancement (34:15), limits of the tech (37:15), what else we can select for (40:00), and unexpected consequences (42:00). DEC 3, 2021
It is a great honor to co-author a paper with Simon Fishel, the last surviving member of the team that produced the first IVF baby (Louise Brown) in 1978. His mentors and collaborators were Robert Edwards (Nobel Prize 2010) and Patrick Steptoe (passed before 2010). ...
Today millions of babies are produced through IVF. In most developed countries roughly 3-5 percent of all births are through IVF, and in Denmark the fraction is about 10 percent! But when the technology was first introduced with the birth of Louise Brown in 1978, the pioneering scientists had to overcome significant resistance.
There may be an alternate universe in which IVF was not allowed to develop, and those millions of children were never born.
Wikipedia: ...During these controversial early years of IVF, Fishel and his colleagues received extensive opposition from critics both outside of and within the medical and scientific communities, including a civil writ for murder.[16] Fishel has since stated that "the whole establishment was outraged" by their early work and that people thought that he was "potentially a mad scientist".[17]
I predict that within 5 years the use of polygenic risk scores will become common in some health systems (i.e., for adults) and in IVF. Reasonable people will wonder why the technology was ever controversial at all, just as in the case of IVF.
Friday, December 03, 2021
Adventures of a Mathematician: Ulam, von Karman, Wiener, and the Golem
[Ulam] ... In Israel many years later, while I was visiting the town of Safed with von Kárman, an old Orthodox Jewish guide with earlocks showed me the tomb of Caro in an old graveyard. When I told him that I was related to a Caro, he fell on his knees... Aunt Caro was directly related to the famous Rabbi Loew of sixteenth-century Prague, who, the legend says, made the Golem — the earthen giant who was protector of the Jews. (Once, when I mentioned this connection with the Golem to Norbert Wiener, he said, alluding to my involvement with Los Alamos and with the H-bomb, "It is still in the family!")
One night in early 1945, just back from Los Alamos, vN woke in a state of alarm in the middle of the night and told his wife Klari:
"... we are creating ... a monster whose influence is going to change history ... this is only the beginning! The energy source which is now being made available will make scientists the most hated and most wanted citizens in any country. The world could be conquered, but this nation of puritans will not grab its chance; we will be able to go into space way beyond the moon if only people could keep pace with what they create ..."
He then predicted the future indispensable role of automation, becoming so agitated that he had to be put to sleep by a strong drink and sleeping pills.
In his obituary for John von Neumann, Ulam recalled a conversation with vN about the
"... ever accelerating progress of technology and changes in the mode of human life, which gives the appearance of approaching some essential singularity in the history of the race beyond which human affairs, as we know them, could not continue."
This is the origin of the concept of technological singularity. Perhaps we can even trace it to that night in 1945 :-)
[p.107] I told Banach about an expression Johnny had used with me in Princeton before stating some non-Jewish mathematician's result, "Die Goim haben den folgenden satz beweisen" (The goys have proved the following theorem). Banach, who was pure goy, thought it was one of the funniest sayings he had ever heard. He was enchanted by its implication that if the goys could do it, then Johnny and I ought to be able to do it better. Johnny did not invent this joke, but he liked it and we started using it.
Saturday, November 27, 2021
Social and Educational Mobility: Denmark vs USA (James Heckman)
Lessons for Americans from Denmark about inequality and social mobility
James Heckman and Rasmus Landersø
Abstract Many progressive American policy analysts point to Denmark as a model welfare state with low levels of income inequality and high levels of income mobility across generations. It has in place many social policies now advocated for adoption in the U.S. Despite generous Danish social policies, family influence on important child outcomes in Denmark is about as strong as it is in the United States. More advantaged families are better able to access, utilize, and influence universally available programs. Purposive sorting by levels of family advantage create neighborhood effects. Powerful forces not easily mitigated by Danish-style welfare state programs operate in both countries.Also discussed in this episode of EconTalk podcast. Russ does not ask the obvious question about disentangling family environment from genetic transmission of inequality.
Sunday, November 14, 2021
Has Hawking's Black Hole Information Paradox Been Resolved?
Do Black Holes Destroy Information?
https://arxiv.org/abs/hep-th/9209058
John Preskill
I review the information loss paradox that was first formulated by Hawking, and discuss possible ways of resolving it. All proposed solutions have serious drawbacks. I conclude that the information loss paradox may well presage a revolution in fundamental physics.
Lessons from the Information Paradox
https://arxiv.org/abs/2012.05770
Suvrat Raju
Abstract: We review recent progress on the information paradox. We explain why exponentially small correlations in the radiation emitted by a black hole are sufficient to resolve the original paradox put forward by Hawking. We then describe a refinement of the paradox that makes essential reference to the black-hole interior. This analysis leads to a broadly-applicable physical principle: in a theory of quantum gravity, a copy of all the information on a Cauchy slice is also available near the boundary of the slice. This principle can be made precise and established — under weak assumptions, and using only low-energy techniques — in asymptotically global AdS and in four dimensional asymptotically flat spacetime. When applied to black holes, this principle tells us that the exterior of the black hole always retains a complete copy of the information in the interior. We show that accounting for this redundancy provides a resolution of the information paradox for evaporating black holes ...
Wednesday, November 10, 2021
Fundamental limit on angular measurements and rotations from quantum mechanics and general relativity (published version)
Physics Letters B Volume 823, 10 December 2021, 136763
Fundamental limit on angular measurements and rotations from quantum mechanics and general relativity
Xavier Calmet and Stephen D.H. Hsu
https://doi.org/10.1016/j.physletb.2021.136763
Abstract
We show that the precision of an angular measurement or rotation (e.g., on the orientation of a qubit or spin state) is limited by fundamental constraints arising from quantum mechanics and general relativity (gravitational collapse). The limiting precision is 1/r in Planck units, where r is the physical extent of the (possibly macroscopic) device used to manipulate the spin state. This fundamental limitation means that spin states cannot be experimentally distinguished from each other if they differ by a sufficiently small rotation. Experiments cannot exclude the possibility that the space of quantum state vectors (i.e., Hilbert space) is fundamentally discrete, rather than continuous. We discuss the implications for finitism: does physics require infinity or a continuum?
In the revision we edited the second paragraph below to clarify the history regarding Hilbert's program, Gödel, and the status of the continuum in analysis. The continuum was quite controversial at the time and was one of the primary motivations for Hilbert's axiomatization. There is a kind of modern middle-brow view that epsilon and delta proofs are sufficient to resolve the question of rigor in analysis, but this ignores far more fundamental problems that forced Hilbert, von Neumann, Weyl, etc. to resort to logic and set theory.
In the early 20th century little was known about neuroscience (i.e., our finite brains made of atoms), and it had not been appreciated that the laws of physics themselves might contain internal constraints that prevent any experimental test of infinitely continuous structures. Hence we can understand Weyl's appeal to human intuition as a basis for the mathematical continuum (Platonism informed by Nature; Gödel was also a kind of Platonist), even if today it appears implausible. Now we suspect that our minds are simply finite machines and nothing more, and that Nature itself does not require a continuum -- i.e., it can be simulated perfectly well with finitary processes.
It may come as a surprise to physicists that infinity and the continuum are even today the subject of debate in mathematics and the philosophy of mathematics. Some mathematicians, called finitists, accept only finite mathematical objects and procedures [30]. The fact that physics does not require infinity or a continuum is an important empirical input to the debate over finitism. For example, a finitist might assert (contra the Platonist perspective adopted by many mathematicians) that human brains built from finite arrangements of atoms, and operating under natural laws (physics) that are finitistic, are unlikely to have trustworthy intuitions concerning abstract concepts such as the continuum. These facts about the brain and about physical laws stand in contrast to intuitive assumptions adopted by many mathematicians. For example, Weyl (Das Kontinuum [26], [27]) argues that our intuitions concerning the continuum originate in the mind's perception of the continuity of space-time.
There was a concerted effort beginning in the 20th century to place infinity and the continuum on a rigorous foundation using logic and set theory. As demonstrated by Gödel, Hilbert's program of axiomatization using finitary methods (originally motivated, in part, by the continuum in analysis) could not succeed. Opinions are divided on modern approaches which are non-finitary. For example, the standard axioms of Zermelo-Fraenkel (ZFC) set theory applied to infinite sets lead to many counterintuitive results such as the Banach-Tarski Paradox: given any two solid objects, the cut pieces of either one can be reassembled into the other [28]. When examined closely all of the axioms of ZFC (e.g., Axiom of Choice) are intuitively obvious if applied to finite sets, with the exception of the Axiom of Infinity, which admits infinite sets. (Infinite sets are inexhaustible, so application of the Axiom of Choice leads to pathological results.) The Continuum Hypothesis, which proposes that there is no cardinality strictly between that of the integers and reals, has been shown to be independent (neither provable nor disprovable) in ZFC [29]. Finitists assert that this illustrates how little control rigorous mathematics has on even the most fundamental properties of the continuum.Weyl was never satisfied that the continuum and classical analysis had been placed on a solid foundation.
Das Kontinuum (Stanford Encyclopedia of Philosophy)
Another mathematical “possible” to which Weyl gave a great deal of thought is the continuum. During the period 1918–1921 he wrestled with the problem of providing the mathematical continuum—the real number line—with a logically sound formulation. Weyl had become increasingly critical of the principles underlying the set-theoretic construction of the mathematical continuum. He had come to believe that the whole set-theoretical approach involved vicious circles[11] to such an extent that, as he says, “every cell (so to speak) of this mighty organism is permeated by contradiction.” In Das Kontinuum he tries to overcome this by providing analysis with a predicative formulation—not, as Russell and Whitehead had attempted, by introducing a hierarchy of logically ramified types, which Weyl seems to have regarded as excessively complicated—but rather by confining the comprehension principle to formulas whose bound variables range over just the initial given entities (numbers). Accordingly he restricts analysis to what can be done in terms of natural numbers with the aid of three basic logical operations, together with the operation of substitution and the process of “iteration”, i.e., primitive recursion. Weyl recognized that the effect of this restriction would be to render unprovable many of the central results of classical analysis—e.g., Dirichlet’s principle that any bounded set of real numbers has a least upper bound[12]—but he was prepared to accept this as part of the price that must be paid for the security of mathematics.
As Weyl saw it, there is an unbridgeable gap between intuitively given continua (e.g. those of space, time and motion) on the one hand, and the “discrete” exact concepts of mathematics (e.g. that of natural number[13]) on the other. The presence of this chasm meant that the construction of the mathematical continuum could not simply be “read off” from intuition. It followed, in Weyl’s view, that the mathematical continuum must be treated as if it were an element of the transcendent realm, and so, in the end, justified in the same way as a physical theory. It was not enough that the mathematical theory be consistent; it must also be reasonable.
Das Kontinuum embodies Weyl’s attempt at formulating a theory of the continuum which satisfies the first, and, as far as possible, the second, of these requirements. In the following passages from this work he acknowledges the difficulty of the task:
… the conceptual world of mathematics is so foreign to what the intuitive continuum presents to us that the demand for coincidence between the two must be dismissed as absurd. (Weyl 1987, 108)
… the continuity given to us immediately by intuition (in the flow of time and of motion) has yet to be grasped mathematically as a totality of discrete “stages” in accordance with that part of its content which can be conceptualized in an exact way.
…See also The History of the Planck Length and the Madness of Crowds.
Tuesday, November 09, 2021
The Balance of Power in the Western Pacific and the Death of the Naval Surface Ship
See LEO SAR, hypersonics, and the death of the naval surface ship:
In an earlier post we described how sea blockade (e.g., against Japan or Taiwan) can be implemented using satellite imaging and missiles, drones, AI/ML. Blue water naval dominance is not required.
PLAN/PLARF can track every container ship and oil tanker as they approach Kaohsiung or Nagoya. All are in missile range -- sitting ducks. Naval convoys will be just as vulnerable.
Sink one tanker or cargo ship, or just issue a strong warning, and no shipping company in the world will be stupid enough to try to run the blockade.
USN guy: We'll just hide the carrier from the satellite and missile seekers using, you know, countermeasures! [Aside: don't cut my carrier budget!]
USAF guy: Uh, the much smaller AESA/IR seeker on their AAM can easily detect an aircraft from much longer ranges. How will you hide a huge ship?
USN guy: We'll just shoot down the maneuvering hypersonic missile using, you know, methods. [Aside: don't cut my carrier budget!]
Missile defense guy: Can you explain to us how to do that? If the incoming missile maneuvers we have to adapt the interceptor trajectory (in real time) to where we project the missile to be after some delay. But we can't know its trajectory ahead of time, unlike for a ballistic (non-maneuvering) warhead.
Monday, November 01, 2021
Preimplantation Genetic Testing for Aneuploidy: New Methods and Higher Pregnancy Rates
Comparison of Outcomes from Concurrent Use of 3 Different PGT-A Laboratories
Oct 18 2021 annual meeting of the American Society for Reproductive Medicine (ASRM)
Klaus Wiemer, PhD
Sunday, October 31, 2021
Demis Hassabis: Using AI to accelerate scientific discovery (protein folding) + Bonus: Bruno Pontecorvo
I want to be remembered as a great physicist, not as your fucking spy!
Saturday, October 30, 2021
Slowed canonical progress in large fields of science (PNAS)
Slowed canonical progress in large fields of science
Johan S. G. Chu and James A. Evans
PNAS October 12, 2021 118 (41) e2021636118
Significance The size of scientific fields may impede the rise of new ideas. Examining 1.8 billion citations among 90 million papers across 241 subjects, we find a deluge of papers does not lead to turnover of central ideas in a field, but rather to ossification of canon. Scholars in fields where many papers are published annually face difficulty getting published, read, and cited unless their work references already widely cited articles. New papers containing potentially important contributions cannot garner field-wide attention through gradual processes of diffusion. These findings suggest fundamental progress may be stymied if quantitative growth of scientific endeavors—in number of scientists, institutes, and papers—is not balanced by structures fostering disruptive scholarship and focusing attention on novel ideas.
Abstract In many academic fields, the number of papers published each year has increased significantly over time. Policy measures aim to increase the quantity of scientists, research funding, and scientific output, which is measured by the number of papers produced. These quantitative metrics determine the career trajectories of scholars and evaluations of academic departments, institutions, and nations. Whether and how these increases in the numbers of scientists and papers translate into advances in knowledge is unclear, however. Here, we first lay out a theoretical argument for why too many papers published each year in a field can lead to stagnation rather than advance. The deluge of new papers may deprive reviewers and readers the cognitive slack required to fully recognize and understand novel ideas. Competition among many new ideas may prevent the gradual accumulation of focused attention on a promising new idea. Then, we show data supporting the predictions of this theory. When the number of papers published per year in a scientific field grows large, citations flow disproportionately to already well-cited papers; the list of most-cited papers ossifies; new papers are unlikely to ever become highly cited, and when they do, it is not through a gradual, cumulative process of attention gathering; and newly published papers become unlikely to disrupt existing work. These findings suggest that the progress of large scientific fields may be slowed, trapped in existing canon. Policy measures shifting how scientific work is produced, disseminated, consumed, and rewarded may be called for to push fields into new, more fertile areas of study.
A toy model of the dynamics of scientific research, with probability distributions for accuracy of experimental results, mechanisms for updating of beliefs by individual scientists, crowd behavior, bounded cognition, etc. can easily exhibit parameter regions where progress is limited (one could even find equilibria in which most beliefs held by individual scientists are false!). Obviously the complexity of the systems under study and the quality of human capital in a particular field are important determinants of the rate of progress and its character.
In physics it is said that successful new theories swallow their predecessors whole. That is, even revolutionary new theories (e.g., special relativity or quantum mechanics) reduce to their predecessors in the previously studied circumstances (e.g., low velocity, macroscopic objects). Swallowing whole is a sign of proper function -- it means the previous generation of scientists was competent: what they believed to be true was (at least approximately) true. Their models were accurate in some limit and could continue to be used when appropriate (e.g., Newtonian mechanics).
In some fields (not to name names!) we don't see this phenomenon. Rather, we see new paradigms which wholly contradict earlier strongly held beliefs that were predominant in the field* -- there was no range of circumstances in which the earlier beliefs were correct. We might even see oscillations of mutually contradictory, widely accepted paradigms over decades.
It takes a serious interest in the history of science (and some brainpower) to determine which of the two regimes above describes a particular area of research. I believe we have good examples of both types in the academy.
* This means the earlier (or later!) generation of scientists in that field was incompetent. One or more of the following must have been true: their experimental observations were shoddy, they derived overly strong beliefs from weak data, they allowed overly strong priors to determine their beliefs.
Blog Archive
Labels
- physics (420)
- genetics (325)
- globalization (301)
- genomics (295)
- technology (282)
- brainpower (280)
- finance (275)
- american society (261)
- China (249)
- innovation (231)
- ai (206)
- economics (202)
- psychometrics (190)
- science (172)
- psychology (169)
- machine learning (166)
- biology (163)
- photos (162)
- genetic engineering (150)
- universities (150)
- travel (144)
- podcasts (143)
- higher education (141)
- startups (139)
- human capital (127)
- geopolitics (124)
- credit crisis (115)
- political correctness (108)
- iq (107)
- quantum mechanics (107)
- cognitive science (103)
- autobiographical (97)
- politics (93)
- careers (90)
- bounded rationality (88)
- social science (86)
- history of science (85)
- realpolitik (85)
- statistics (83)
- elitism (81)
- talks (80)
- evolution (79)
- credit crunch (78)
- biotech (76)
- genius (76)
- gilded age (73)
- income inequality (73)
- caltech (68)
- books (64)
- academia (62)
- history (61)
- intellectual history (61)
- MSU (60)
- sci fi (60)
- harvard (58)
- silicon valley (58)
- mma (57)
- mathematics (55)
- education (53)
- video (52)
- kids (51)
- bgi (48)
- black holes (48)
- cdo (45)
- derivatives (43)
- neuroscience (43)
- affirmative action (42)
- behavioral economics (42)
- economic history (42)
- literature (42)
- nuclear weapons (42)
- computing (41)
- jiujitsu (41)
- physical training (40)
- film (39)
- many worlds (39)
- quantum field theory (39)
- expert prediction (37)
- ufc (37)
- bjj (36)
- bubbles (36)
- mortgages (36)
- google (35)
- race relations (35)
- hedge funds (34)
- security (34)
- von Neumann (34)
- meritocracy (31)
- feynman (30)
- quants (30)
- taiwan (30)
- efficient markets (29)
- foo camp (29)
- movies (29)
- sports (29)
- music (28)
- singularity (27)
- entrepreneurs (26)
- conferences (25)
- housing (25)
- obama (25)
- subprime (25)
- venture capital (25)
- berkeley (24)
- epidemics (24)
- war (24)
- wall street (23)
- athletics (22)
- russia (22)
- ultimate fighting (22)
- cds (20)
- internet (20)
- new yorker (20)
- blogging (19)
- japan (19)
- scifoo (19)
- christmas (18)
- dna (18)
- gender (18)
- goldman sachs (18)
- university of oregon (18)
- cold war (17)
- cryptography (17)
- freeman dyson (17)
- smpy (17)
- treasury bailout (17)
- algorithms (16)
- autism (16)
- personality (16)
- privacy (16)
- Fermi problems (15)
- cosmology (15)
- happiness (15)
- height (15)
- india (15)
- oppenheimer (15)
- probability (15)
- social networks (15)
- wwii (15)
- fitness (14)
- government (14)
- les grandes ecoles (14)
- neanderthals (14)
- quantum computers (14)
- blade runner (13)
- chess (13)
- hedonic treadmill (13)
- nsa (13)
- philosophy of mind (13)
- research (13)
- aspergers (12)
- climate change (12)
- harvard society of fellows (12)
- malcolm gladwell (12)
- net worth (12)
- nobel prize (12)
- pseudoscience (12)
- Einstein (11)
- art (11)
- democracy (11)
- entropy (11)
- geeks (11)
- string theory (11)
- television (11)
- Go (10)
- ability (10)
- complexity (10)
- dating (10)
- energy (10)
- football (10)
- france (10)
- italy (10)
- mutants (10)
- nerds (10)
- olympics (10)
- pop culture (10)
- crossfit (9)
- encryption (9)
- eugene (9)
- flynn effect (9)
- james salter (9)
- simulation (9)
- tail risk (9)
- turing test (9)
- alan turing (8)
- alpha (8)
- ashkenazim (8)
- data mining (8)
- determinism (8)
- environmentalism (8)
- games (8)
- keynes (8)
- manhattan (8)
- new york times (8)
- pca (8)
- philip k. dick (8)
- qcd (8)
- real estate (8)
- robot genius (8)
- success (8)
- usain bolt (8)
- Iran (7)
- aig (7)
- basketball (7)
- free will (7)
- fx (7)
- game theory (7)
- hugh everett (7)
- inequality (7)
- information theory (7)
- iraq war (7)
- markets (7)
- paris (7)
- patents (7)
- poker (7)
- teaching (7)
- vietnam war (7)
- volatility (7)
- anthropic principle (6)
- bayes (6)
- class (6)
- drones (6)
- econtalk (6)
- empire (6)
- global warming (6)
- godel (6)
- intellectual property (6)
- nassim taleb (6)
- noam chomsky (6)
- prostitution (6)
- rationality (6)
- academia sinica (5)
- bobby fischer (5)
- demographics (5)
- fake alpha (5)
- kasparov (5)
- luck (5)
- nonlinearity (5)
- perimeter institute (5)
- renaissance technologies (5)
- sad but true (5)
- software development (5)
- solar energy (5)
- warren buffet (5)
- 100m (4)
- Poincare (4)
- assortative mating (4)
- bill gates (4)
- borges (4)
- cambridge uk (4)
- censorship (4)
- charles darwin (4)
- computers (4)
- creativity (4)
- hormones (4)
- humor (4)
- judo (4)
- kerviel (4)
- microsoft (4)
- mixed martial arts (4)
- monsters (4)
- moore's law (4)
- soros (4)
- supercomputers (4)
- trento (4)
- 200m (3)
- babies (3)
- brain drain (3)
- charlie munger (3)
- cheng ting hsu (3)
- chet baker (3)
- correlation (3)
- ecosystems (3)
- equity risk premium (3)
- facebook (3)
- fannie (3)
- feminism (3)
- fst (3)
- intellectual ventures (3)
- jim simons (3)
- language (3)
- lee kwan yew (3)
- lewontin fallacy (3)
- lhc (3)
- magic (3)
- michael lewis (3)
- mit (3)
- nathan myhrvold (3)
- neal stephenson (3)
- olympiads (3)
- path integrals (3)
- risk preference (3)
- search (3)
- sec (3)
- sivs (3)
- society generale (3)
- systemic risk (3)
- thailand (3)
- twitter (3)
- alibaba (2)
- bear stearns (2)
- bruce springsteen (2)
- charles babbage (2)
- cloning (2)
- david mamet (2)
- digital books (2)
- donald mackenzie (2)
- drugs (2)
- dune (2)
- exchange rates (2)
- frauds (2)
- freddie (2)
- gaussian copula (2)
- heinlein (2)
- industrial revolution (2)
- james watson (2)
- ltcm (2)
- mating (2)
- mba (2)
- mccain (2)
- monkeys (2)
- national character (2)
- nicholas metropolis (2)
- no holds barred (2)
- offices (2)
- oligarchs (2)
- palin (2)
- population structure (2)
- prisoner's dilemma (2)
- singapore (2)
- skidelsky (2)
- socgen (2)
- sprints (2)
- star wars (2)
- ussr (2)
- variance (2)
- virtual reality (2)
- war nerd (2)
- abx (1)
- anathem (1)
- andrew lo (1)
- antikythera mechanism (1)
- athens (1)
- atlas shrugged (1)
- ayn rand (1)
- bay area (1)
- beats (1)
- book search (1)
- bunnie huang (1)
- car dealers (1)
- carlos slim (1)
- catastrophe bonds (1)
- cdos (1)
- ces 2008 (1)
- chance (1)
- children (1)
- cochran-harpending (1)
- cpi (1)
- david x. li (1)
- dick cavett (1)
- dolomites (1)
- eharmony (1)
- eliot spitzer (1)
- escorts (1)
- faces (1)
- fads (1)
- favorite posts (1)
- fiber optic cable (1)
- francis crick (1)
- gary brecher (1)
- gizmos (1)
- greece (1)
- greenspan (1)
- hypocrisy (1)
- igon value (1)
- iit (1)
- inflation (1)
- information asymmetry (1)
- iphone (1)
- jack kerouac (1)
- jaynes (1)
- jazz (1)
- jfk (1)
- john dolan (1)
- john kerry (1)
- john paulson (1)
- john searle (1)
- john tierney (1)
- jonathan littell (1)
- las vegas (1)
- lawyers (1)
- lehman auction (1)
- les bienveillantes (1)
- lowell wood (1)
- lse (1)
- machine (1)
- mcgeorge bundy (1)
- mexico (1)
- michael jackson (1)
- mickey rourke (1)
- migration (1)
- money:tech (1)
- myron scholes (1)
- netwon institute (1)
- networks (1)
- newton institute (1)
- nfl (1)
- oliver stone (1)
- phil gramm (1)
- philanthropy (1)
- philip greenspun (1)
- portfolio theory (1)
- power laws (1)
- pyschology (1)
- randomness (1)
- recession (1)
- sales (1)
- skype (1)
- standard deviation (1)
- starship troopers (1)
- students today (1)
- teleportation (1)
- tierney lab blog (1)
- tomonaga (1)
- tyler cowen (1)
- venice (1)
- violence (1)
- virtual meetings (1)
- wealth effect (1)