Bloomberg: Simone Collins knew she was pregnant the moment she answered the phone. ... Embryo 3, the fertilized egg that Collins and her husband, Malcolm, had picked, could soon be their daughter—a little girl with, according to their tests, an unusually good chance of avoiding heart disease, cancer, diabetes, and schizophrenia.
This isn’t a story about Gattaca-style designer babies. No genes were edited in the creation of Collins’s embryo. The promise, from dozens of fertility clinics around the world, is just that the new DNA tests they’re using can assess, in unprecedented detail, whether one embryo is more likely than the next to develop a range of illnesses long thought to be beyond DNA-based predictions. It’s a new twist on the industry-standard testing known as preimplantation genetic testing, which for decades has checked embryos for rare diseases, such as cystic fibrosis, that are caused by a single gene.
One challenge with leading killers like cancer and heart disease is that they’re usually polygenic: linked to many different genes with complex interactions. Patients such as Collins can now take tests that assess thousands of DNA data points to decode these complexities and compute the disease risks. Genomic Prediction, the five-year-old New Jersey company that handled the tests for her fertility clinic, generates polygenic risk scores, predicting in percentage terms each embryo’s chances of contracting each disease in the panel, plus a composite score for overall health. Parents with multiple embryos can then weigh the scores when deciding which one to implant.
...
This new form of genetic embryo testing appears to move humanity one step closer to control of its evolution. The $14 billion IVF industry brings more than 500,000 babies into the world each year, and with infertility rates rising, the market is expected to more than double this decade. Companies including Genomic Prediction bet many going into that process have seen enough loved ones suffer from a polygenic disease to want risk scoring.
[ Note I think the number of IVF babies born worldwide each year is more like 1 million, but there is some uncertainty in estimates. ]
...
In December, Genomic Prediction doubled its venture funding to about $25 million and says it will use the cash to expand and add to its testing panel. Boston IVF, one of the biggest fertility networks in the US, recently started offering Genomic Prediction’s polygenic testing to its patients, says CEO David Stern. “Like anything else, you have early adopters,” he says. “We have had patients who worked in the biotech field or the Harvard milieu who came in and asked for it.” Stern predicts that, like egg freezing, polygenic embryo testing will grow slowly at first, but steadily, and eventually demand will reflect the powerful appeal of lowering a child’s odds for disease.
...
Believers such as Collins and her husband support government subsidies for fertility and parenthood but aren’t interested in any conversation about slowing down. “This is about the people who care about giving their children every opportunity,” she says. “I do not believe that law or social norms are going to stop parents from giving their kids advantages.”
An important point that is not discussed in the article (understandable, given the complexity of the topics listed above), is that precise genotyping of embryos leads to higher success rates in IVF.
... improved success rates resulting from higher accuracy in aneuploidy screening of embryos will affect millions of families around the world, and over 60% of all IVF families in the US.
The SNP array platform allows very accurate genotyping of each embryo at ~1 million locations in the genome, and the subsequent bioinformatic analysis produces a much more accurate prediction of chromosomal normality than the older methods.
Millions of embryos are screened each year using PGT-A, about 60% of all IVF embryos in the US.
Klaus Wiemer is the laborator director for Poma Fertility near Seattle. He conducted this study independently, without informing Genomic Prediction.
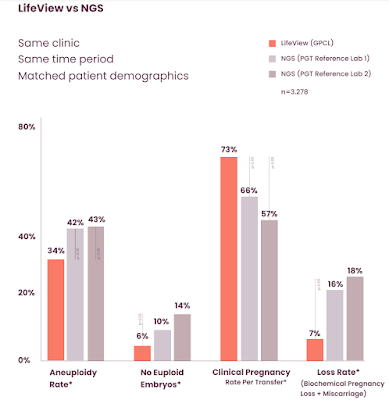
There are ~3000 embryos in the dataset, all biopsied at Poma and samples allocated to three testing labs A,B,C using the two different methods. The family demographics (e.g., maternal age) were similar in all three groups. Lab B is Genomic Prediction and A,C are two of the largest IVF testing labs in the world, using NGS.
The results imply lower false-positive rates, lower false-negative rates, and higher accuracy overall from our methods. These lead to a significantly higher pregnancy success rate.
The new technology has the potential to help millions of families all over the world.
Prioritizing research in IVF using large sample sizes could plausibly raise success rates per cycle to, e.g., ~80%. The qualitative experience of parents using IVF will improve with average success rates, perhaps relieving much of the angst and uncertainty.


No comments:
Post a Comment